About me
In July 2018 I finished a PhD in statistics in Duke's Department of Statistical Science, advised by Scott Schmidler. I received my Bachelor of Science degree in 2006 from University of Richmond, studying math and a bit of physics and biology.
Research
Graduate
My dissertation projects were focused on improving statistical modeling of protein structure evolution and alignment. In one project I have investigated how to account for dependence structure in structure evolution - most existing methods assume independence for simplicity (computational and otherwise), but this assumption is certainly violated in reality. How do dependent models perform differently from independent models? What can be gained from incorporating dependence? Our work demonstrated that ignoring dependence in structural evolution can bias evolutionary inference and affect phylogenetic inference. This work was presented at the International Conference for Research in Computational Biology in April 2018. The abstract can be viewed here.
In a second project my advisor and I have developed a model extension that allows evolutionary modeling of flexible proteins. This manuscript is in preparation. In the graphic below, the figure at left shows a pair of conformations of a protein with a flexible "lid" region that opens or closes based on ligand binding.
My dissertation projects were focused on improving statistical modeling of protein structure evolution and alignment. In one project I have investigated how to account for dependence structure in structure evolution - most existing methods assume independence for simplicity (computational and otherwise), but this assumption is certainly violated in reality. How do dependent models perform differently from independent models? What can be gained from incorporating dependence? Our work demonstrated that ignoring dependence in structural evolution can bias evolutionary inference and affect phylogenetic inference. This work was presented at the International Conference for Research in Computational Biology in April 2018. The abstract can be viewed here.
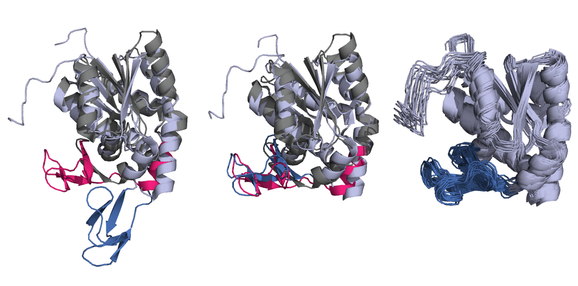
In a second project my advisor and I have developed a model extension that allows evolutionary modeling of flexible proteins. This manuscript is in preparation. In the graphic below, the figure at left shows a pair of conformations of a protein with a flexible "lid" region that opens or closes based on ligand binding.
Structural models that don't accommodate flexibility will use a single, rigid-body transformation to superimpose these two structures, and can only align them as well as the left-most figure: the "lid" region is poorly aligned as it is "open" in one structure (dark blue) and "closed" in the other (pink). In the middle figure, our flexible structural model has identified the correct positions of the flexion points which delineate the flexible lid region and is able to align all positions well. The right-most figure visualizes the Bayesian posterior distribution of the parameters used to transform (rotate, translate, etc.) the proteins' coordinates close to each other: in this case, the batch of twenty samples of the rotated protein show that the posterior distribution of the model's registration parameters has high posterior precision.
For computer code, see https://github.com/garylarson/SEMAP. If you can't reach the repo at that link, it probably means I'm still working on user-friendliness of the repo and it's still private. In that case, contact me via email for access.
For computer code, see https://github.com/garylarson/SEMAP. If you can't reach the repo at that link, it probably means I'm still working on user-friendliness of the repo and it's still private. In that case, contact me via email for access.
Undergraduate
At University of Richmond I worked with Ted Bunn on a project related to the cosmic microwave background (CMB) radiation using data from the WMAP satellite. In our research we tried to reliably improve low-level galactic dust identification/removal techniques by working with wavelet or radon transformations of the CMB data. In a broad sense, we were investigating a potential counterexample of an implication made by the prevailing cosmological model of the universe.
At University of Richmond I worked with Ted Bunn on a project related to the cosmic microwave background (CMB) radiation using data from the WMAP satellite. In our research we tried to reliably improve low-level galactic dust identification/removal techniques by working with wavelet or radon transformations of the CMB data. In a broad sense, we were investigating a potential counterexample of an implication made by the prevailing cosmological model of the universe.